

133 Molesworth Street
PO Box 5013
Wellington 6140
New Zealand
T+64 4 496 2000
14 December 2022
James
By email: [FYI request #21306 email]
Ref: H2022018035
Tēnā koe James
Response to your request for official information
Thank you for your request under the Official Information Act 1982 (the Act) transferred to
Manatū Hauora (the Ministry of Health) from Te Whatu Ora- Health New Zealand (Te Whatu
Ora) on 2 December 2022 for information regarding COVID-19 infectious period calculations
and processes. You requested:
“I am seeking the information and directives given to the National Contact Tracing Service
and any other COVID-19 officials around infectious period calculations and processes.
Specifically, I am looking for the directives that relate to when a person's day 0 and
infectious period begins when they have tested positive more than 7 days after symptom
onset. I am also looking for the health advice that supports such directives.”
One document titled
Efficacy of Test to Release has been identified within scope of your
request. This document is attached to this letter as Appendix 1 and is released to you in full.
I trust this information fulfils your request. Under section 28(3) of the Act, you have the right to
ask the Ombudsman to review any decisions made under this request. The Ombudsman may
be contacted by email at:
[email address] or by calling 0800 802 602.
Please note that this response, with your personal details removed, may be published on the
Manatū Hauora website at:
www.health.govt.nz/about-ministry/information-releases/responses-
official-information-act-requests.
Nāku noa, nā
Dave Henderson
Interim Group Manager, Intelligence, Surveillance and Knowledge
Public Health Agency | Te Pou Hauora Tūmatanui
Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
Efficacy of Test to Release
The purpose of this document is to provide evidence for the efficacy of Rapid Antigen Tests
(RATs) in a “Test to Release” schedule.
The current period of isolation is a fixed period of 7 days from the date of onset of symptoms, or
diagnosis, whichever is earlier. A test to release schedule uses a combination of a lower and upper
fixed period of time for isolation combined with release form isolation if an individual tests negative
between these dates. The rationale is that RATs are able to predict infectiousness with sufficient
accuracy to ensure infectious individuals are isolated, while non-infectious individuals are released
from isolation.
The ability of RATs to identify infectious individuals in a “Test to Release Schedule” does not need to
be 100% accurate for this method to be used to determine the period of isolation. Instead, the
ACT 1982
evidence should provide sufficient basis to conclude that a “Test to Release Schedule” is superior to
a “Fixed Isolation Period” of 7 (or some other number of) days. In this setting, superiority would be
inferred, if there was evidence that a “Test to Release” schedule resulted in fewer infectious
individuals being released into the community without a significant increase in non-infectious
individuals being isolated or a similar proportion of individuals being released with a significantly
deceased period of isolation in non-infectious individuals.
Summary.
INFORMATION
• The performance of test to release strategy primarily depends upon the ability of rapid
antigen tests to differentiate between those who are infectious and those who are not
infectious. As the cohort of individuals isolating have already been diagnosed using RATs, the
subsequent test performance is expected to be high, as those who have a false negative RAT
will not be captured by either strategy.
• Infectiousness is not directly measurable. The current most reliable measure of
infectiousness is the ability to culture live virus. Culture of virus is not possible to use in a
clinical setting as the tests are too expensive and time consuming to use on a large scale. An
individual who is “culture negative” is very unlikely to be infectious. An individual who is
“culture positive” is potentially infectious.
• It is well recognised that infectiousness is not evenly distributed throughout those
individuals who test positive. Both biological and behavioural factors will influence this
variation in infectiousness. Some individuals are substantially more infectious than others. It
has been estimated that the majority of transmission occurs from a minority of individuals.
This observation indicates that individuals who are highly infectious are also highly likely to
return a positive RAT test.
• It is well recognised that infectiousness varies marke
RELEASED UNDER THE OFFICIAL dly over time. As RATs measure viral
antigens, not intact virions, it is likely that the relationship between a positive RAT and
infectiousness will also decrease over time.
•
The current evidence would support the assumption that within the first week of
infection, a positive RAT is strongly correlated with culturable virus and that the individual
is infectious. However, a negative RAT early in the course of disease (before day 5) does
not guarantee an individual wil not be infectious.
•
The current evidence would support the assumption that after the first week of infection,
a negative RAT is strongly correlated with non-culturable virus and that the individual is
1


Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
unlikely to be infectious. A positive RAT is correlated with infectiousness, but not as
strongly as within the first week. At more than 14 days either a positive RAT or culturable
virus are uncommon.
• The transmissibility of Omicron variant is markedly increased compared to the original
Wuhan or later Delta variants. Much of this is due to immune evasion, but there is also an
element of increased infectiousness. Data regarding the rates or risks of infectiousness from
previous variants must be interpreted with caution for the current outbreak.
The relationship between PCR Ct value and culture positivity
There is a clear relationship between the Ct value and Culture positivity. Culture positivity decreases
as the Ct value rises, which is assumed to be due to a decreasing viral load. Virus is almost always
culturable at a Ct value of 25 or less, decreasing to less than 10% at a Ct value of 35 or more (Figs
1&2) (1) (2). However, there is a stronger relationship between culture positivity and time since the
beginning of infection, indicating that the relationship between CT value and culture positivity will
vary over the course of an infection.
ACT 1982
Fig 1. Relationship between RT-PCR Ct value time since infection and culture positivity.
INFORMATION
Fig 2 Relationship between culture and PCR Ct values
RELEASED UNDER THE OFFICIAL
2
Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
Omicron
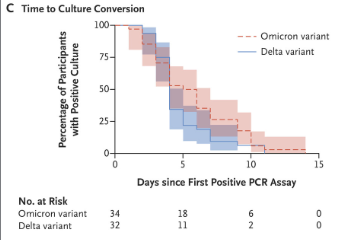
Are more recent study analysing differences in the duration of shedding of the Omicron and Delta
variants reported that the time to culture conversion was greater for Omicron than Delta (3). For
Omicron, the proportion that are still shedding culturable virus at day 10 was 25%, while for Delta,
the proportion was 6%. Culturable virus was not recovered after day 14.
ACT 1982
The relationship between RATs, PCR Ct and positive cultures
The Human Challenge Trial assessed the relationship between infection and culturable virus and
reported that the RAT sensitivity for culturable virus exceeded 80%, once a positive PCR test had
INFORMATION
been obtained. The study also reported that a negative RAT was a reliable indicator of a negative
viral culture (4).
A systematic review of the performance of Rapid Antigen Tests (RATs) reported that assays shown to
meet appropriate criteria, such as WHO's priority target product profiles for COVID-19 diagnostics
(‘acceptable’ sensitivity ≥ 80% and specificity ≥ 97%), can be considered as a replacement for
laboratory-based RT-PCR when immediate decisions about patient care must be made, or where RT-
PCR cannot be delivered in a timely manner (5). However, this review also states that “Test accuracy
studies cannot adequately assess the ability of antigen tests to differentiate those who are infectious
and require isolation from those who pose no risk, as there is no reference standard for
infectiousness”. A review of the performance of 14 RATs reported substantial variability in the limit
of detection measured against the Ct value of paired samples, from 26.8 to 34.7. This encompasses
the range of results which occur for many individuals over the entire course of an illness. However,
the most effective RATs demonstrated a true positive rate compared to paired samples from PCR for
values of 99.1% for a Ct value of ≤30 and 90.9% for a Ct value of ≤33 (6).
Al RATs used in New Zealand have undergone a rigorous assessment to ensure that the test has a
sensitivity of at least 80% overall and >90% for CT values <25. RATs are also assessed for usability
RELEASED UNDER THE OFFICIAL
which has been uniformly high.
Similar results have been reported by other studies (3, 7, 8).
The relationship between Culture and Infectiousness
The ability to culture virus does not automatically indicate infectiousness. The minimum infectious
dose for Omicron, or any other variant is still unknown, but the higher the viral load of intact virus,
the greater the risk of infection. The relationship between dose and infection wil be sigmoid, with a
3
link to page 5
Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
very low rate of infection at low doses, rising to a very high rate of infection as the dose increases. It
has been estimated that a sample with 108 RNA copies per ml, a positive culture rate of
approximately 50% will be achieved (9).
Therefore, it is clear that not all individuals from whom culturable virus can be obtained will produce
enough virus to infect other in the majority of exposure events. Indeed an argument can be made
that as the majority of infections are caused by a minority of highly infectious individuals. Two
epidemiological parameters often characterise the transmissibility of infectious diseases: the basic
reproductive number (R0) and the dispersion parameter (k). R0 describes, on average, how many
individuals in a susceptible population will be infected by someone with that disease, and k details
the variation in individual infectiousness. The smaller the k value, the greater the variation. That is,
fewer cases cause the majority of infections, and a greater proportion of infections tend to be linked
to large clusters via superspreading events. (10) During the COVID-19 pandemic, transmission of
SARS-CoV-2 has been highly overdispersed, as 60–75% of cases infect no one and, propelled by
superspreading events, 10–20% of cases cause 80% of secondary infections (11-13)
ACT 1982
Modelling studies have reported that the number of super-emitters of SARS-CoV-2 has increased
progressively so that for the WT, one in 1,000 infected persons was a super-emitter; for Delta one in
30; and for Omicron one in 20 or one in 10, depending on the viral load estimate used
1 . The
infectivity-strengthening mutations N440K, T478K, and N501Y enhance infectiousness. Among them,
T478K is one of two RBD mutations in the Delta variant, while N501Y is presented on many
prevailing variants (14).
INFORMATION
The conclusion is that it is not necessary to identify all of the individuals who are infectious to have
an impact on the rate of transmission, but to identify those who are superspreaders, who are most
likely to have the highest viral load and be RAT positive.
Real world studies of test to release
Although several countries or States have implemented test to release policies, there is no reliable
analysis of the success of these policies. Changes in regulations have often comprised a package of
alterations to interventions, which, in addition to the natural variation in case numbers, results in
difficulty in ascribing a causal relationship to changes in the duration of isolation.
Model ing studies of test to release
Modelling of test to release have been published. Two of the key components of the model include
the sensitivity of the RAT test in predicting infectiousness and the distribution of infectious cases
over time (3). For a RAT sensitivity of between 0.7 and 0.8 and a model which predicts a 16% risk of
infectiousness at day 7, an isolation schedule of at least 7 days would result in 15.8% of released
individuals being infectious for a mean excess isolation per person of 76.8 hours. A test to release
schedule would reduce the proportion of infectious cases released to 9.2% for no significant change
in the mean excess isolation per person of 79.2 hours.
RELEASED UNDER THE OFFICIAL
A model assessing the ability of two consecutive day negative RATs reported that the number of
infectious days in the community can be reduced to almost zero (15). The model was based on data
relating viral load to test positivity over time so maybe less dependent on assumptions about test
performance at different points in time. Testing was just as efficient if commenced on day 3 or day 5.
1D
OI: https://doi.org/10.4414/smw.2022.w30133
Publication Date: 06.01.2022
Swiss Med Wkly. 2022;152:w30133
4
Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
This model assumed that the infection kinetics for Omicron are similar to those for pre-Omicron
variants.
Conclusion
The aim of isolation is to decrease the risk of individuals who are infectious, being released from
isolation. However, infectiousness is not a binary (yes or no) state and there is ample evidence to
support the observations that individuals who are highly infectious, are the primary drivers of
community spread. Therefore the identification of infectiousness does not need to be perfect but to
identify those who are the most infectious. These individuals are likely to within the cohort
identified by a positive RAT and to remain positive until the viral load has substantially reduced. This
time will be variable and for the most infectious likely to be more than 7 days. RATs may be
unreliable at less than 5 days after infection, and be unnecessary more than 10 days after diagnosis
or symptom onset.
ACT 1982
References
1.
Singanayagam A, Patel M, Charlett A, Lopez Bernal J, Saliba V, Ellis J, et al.
Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases
of COVID-19, England, January to May 2020. Euro Surveil . 2020;25(32).
2.
Jefferson T, Spencer EA, Brassey J, Heneghan C. Viral Cultures for Coronavirus
INFORMATION
Disease 2019 Infectivity Assessment: A Systematic Review. Clinical Infectious Diseases.
2020;73(11):e3884-e99.
3.
Bays D, Whiteley T, Pindar M, Taylor J, Walker B, Wil iams H, et al. Mitigating
isolation: The use of rapid antigen testing to reduce the impact of self-isolation
periods. medRxiv. 2021:2021.12.23.21268326.
4.
Kil ingley B, Mann AJ, Kalinova M, Boyers A, Goonawardane N, Zhou J, et al.
Safety, tolerability and viral kinetics during SARS-CoV-2 human chal enge in young
adults. Nature Medicine. 2022.
5.
Dinnes J, Deeks JJ, Berhane S, Taylor M, Adriano A, Davenport C, et al. Rapid,
point-of-care antigen and molecular-based tests for diagnosis of SARS-CoV-2 infection.
Cochrane Database of Systematic Reviews. 2021(3).
6.
Routsias JG, Mavrouli M, Tsoplou P, Dioikitopoulou K, Tsakris A. Diagnostic
performance of rapid antigen tests (RATs) for SARS-CoV-2 and their efficacy in
monitoring the infectiousness of COVID-19 patients. Scientific Reports.
2021;11(1):22863.
7.
Boucau J, Marino C, Regan J, Uddin R, Choudhary MC, Flynn JP, et al. Duration
RELEASED UNDER THE OFFICIAL
of viable virus shedding in SARS-CoV-2 omicron variant infection. medRxiv.
2022:2022.03.01.22271582.
8.
National Institute of Infectious Disease. Active epidemiological investigation on
SARS-CoV-2 infection caused by Omicron variant (Pango lineage B.1.1.529) in Japan:
preliminary report on infectious period. National Institute of Infectious Disease,; 2022
5 Jan 2022.
5
Document 1
TEST TO RELEASE: AN ALTERNATIVE TO FIXED ISOLATION PERIOD.
9.
Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Mül er MA, et al.
Virological assessment of hospitalized patients with COVID-2019. Nature.
2020;581(7809):465-9.
10.
Chen PZ, Koopmans M, Fisman DN, Gu FX. Understanding why superspreading
drives the COVID-19 pandemic but not the H1N1 pandemic. The Lancet Infectious
Diseases. 2021;21(9):1203-4.
11.
Sun K, Wang W, Gao L, Wang Y, Luo K, Ren L, et al. Transmission
heterogeneities, kinetics, and control ability of SARS-CoV-2. Science. 2021;371(6526).
12.
Lau MS, Grenfel B, Thomas M, Bryan M, Nelson K, Lopman B. Characterizing
superspreading events and age-specific infectiousness of SARS-CoV-2 transmission in
Georgia, USA. Proceedings of the National Academy of Sciences. 2020;117(36):22430-
5.
13.
Hasan A, Susanto H, Kasim MF, Nuraini N, Lestari B, Triany D, et al.
ACT 1982
Superspreading in early transmissions of COVID-19 in Indonesia. Scientific reports.
2020;10(1):1-4.
14.
Chen J, Wang R, Gilby NB, Wei G-W. Omicron Variant (B.1.1.529): Infectivity,
Vaccine Breakthrough, and Antibody Resistance. Journal of Chemical Information and
Modeling. 2022;62(2):412-22.
15.
Quilty BJ, Pul iam JRC, Pearson CAB. Test to release from isolation after testing
positive for SARS-CoV-2. medRxiv. 2022:2022.01.04.21268372.
INFORMATION
RELEASED UNDER THE OFFICIAL
6
Document Outline




